LINE-1 retrotransposons are repetitive DNA elements present throughout the genome. Although most of them are truncated and incapable of transferring to new locations, a fraction of them remain active and capable of relocating to new regions of the genome through retrotransposition, a process that can disrupt other genes and sometimes drive tumorigenesis. Zumalave et al. used long-read sequencing to study 10 tumors with unusually high retrotransposition activity and examined the patterns of genetic changes and mechanisms involved in the process of tumor evolution resulting from retrotransposons (see the Perspective by Smits and Richardson). The authors then validated their findings in a lower-retrotransposition cohort, confirming their generalizability. —Yevgeniya Nusinovich
LINE-1 (L1) retrotransposons are abundant repetitive elements that make up ~17% of the human reference genome. These sequences behave as intragenomic parasites, propagating through a “copy-and-paste” mechanism known as retrotransposition. In this process, an active L1 locus is transcribed into an RNA intermediate, which is then reverse-transcribed and inserted at a new genomic site. Although most L1 copies are inactive, a handful of loci generate new insertions. Such mobilization can disrupt genome integrity, promote insertional mutagenesis, and occasionally contribute to human disease, including cancer. In certain tumors, somatic L1 activity becomes a prominent source of mutational burden.
Previous work revealed that aberrant somatic L1 retrotransposition can mediate diverse genomic rearrangements, with deletions representing the most frequent class (>95%). However, these analyses were largely hampered by the limitations of short-read sequencing, which obscures other forms of L1-driven chromosomal instability. In this work, we integrated multiplatform sequencing approaches, with an emphasis on long-read data, to investigate somatic retrotransposition in 10 human tumors selected for their high L1 activity. We developed a bioinformatics algorithm to uncover the patterns and mechanisms of retrotransposon-mediated rearrangements at nucleotide-level resolution. Furthermore, we implemented timing estimation approaches to characterize the tempo of these rearrangements during tumorigenesis.
Across the 10 tumors, we identified 6418 somatic L1 retrotransposition events. Among these, 152 insertions bridged genomic rearrangements. Although deletions remained the dominant category, translocations emerged as the second most frequent type, accounting for ~30% of these forms of structural variants. Notably, approximately half of the retrotransposon bridges mediating interchromosomal junctions participated in reciprocal translocations—balanced exchanges between nonhomologous chromosomes that were largely undetectable in earlier short-read studies. Long-read sequencing enabled reconstruction of the internal architecture of the insertions and revealed that these reciprocal translocations most commonly involve two distinct L1 molecules forming the junctions of the two derivative chromosomes. Notably, we found no evidence for postintegration homologous recombination between preexisting somatic L1 insertions. Instead, our data support a model in which two independent L1 insertions are resolved concurrently to generate the rearrangement. Similar principles also underlie the formation of reciprocal inversions and more complex structural variants, in which independent retrotransposition events mediate multiple junctions across concatenated rearrangements.
Timing analyses revealed that ~65% of the L1 insertion events occurred early during tumor evolution. Comparative analyses in an independent set of tumors spanning lower retrotransposition rates confirmed associations between retrotransposition, TP53 alterations, and global DNA hypomethylation while further implicating disruption of the Fanconi anemia pathway in elevated rearrangement burdens.
Together, our observations establish somatic L1 activity as a major contributor to tumor genome plasticity, extending its impact far beyond classical insertional mutagenesis. Approximately three-quarters of the retrotransposon-mediated rearrangements identified in this work would remain undetected or misclassified by short-read sequencing, which underscores how this class of structural variation has been largely overlooked. Collectively, our findings position L1 activity as an early driver of large-scale chromosomal instability in human cancer, with approximately one in every 40 to 60 somatic insertions giving rise to a genomic rearrangement.
LINE-1 (L1) retrotransposition generates somatic genomic variation in human cancer, but short-read sequencing has limited our understanding of its structural consequences and dynamics. Using long-read sequencing, we analyzed 10 tumors with exceptionally high retrotransposition activity, comprising more than 6000 somatic events. We reveal that L1-mediated reciprocal translocations occur frequently, typically driven by two concurrent L1 retrotransposition events on nonhomologous chromosomes. Using an independent tumor cohort spanning low to high L1 activity, we estimate that retrotransposon-mediated rearrangements arise at a frequency of one event per 60 somatic retrotranspositions. Molecular timing analyses indicate that these events arise early in tumorigenesis, establishing L1 activity as an early driver of chromosomal instability. Our findings demonstrate that L1 contributes substantially to cancer genome evolution in certain tumors.
LINE-1逆转录转座子是遍布基因组的重复DNA元件。尽管其中大多数已被截短且无法转移到新位置,但仍有部分保持活性,能够通过逆转录转座过程重新定位到基因组的新区域,这一过程可能破坏其他基因,有时还会驱动肿瘤发生。Zumalave等人使用长读长测序技术研究了10个具有异常高逆转录转座活性的肿瘤,并分析了由逆转录转座子导致的遗传变化模式及肿瘤演化过程中所涉及的机制(参见Smits和Richardson的观点文章)。随后,作者在一个逆转录转座活性较低的队列中验证了他们的发现,证实了其普遍性。——Yevgeniya Nusinovich
LINE-1 (L1) 逆转录转座子是丰富的重复元件,约占人类参考基因组的17%。这些序列作为基因组内寄生生物,通过一种称为逆转录转座的“复制-粘贴”机制进行增殖。在这个过程中,一个活跃的L1位点被转录成RNA中间体,然后被逆转录并插入到新的基因组位点。尽管大多数L1拷贝是无活性的,但少数位点会产生新的插入。这种活动会破坏基因组完整性,促进插入突变,并偶尔导致人类疾病,包括癌症。在某些肿瘤中,体细胞L1活性成为突变负荷的一个重要来源。
先前的研究表明,异常的体细胞L1逆转录转座可以介导多种基因组重排,其中缺失是最常见的类型(>95%)。然而,这些分析很大程度上受到短读长测序局限性的阻碍,这种技术掩盖了其他形式的L1驱动的染色体不稳定性。在这项工作中,我们整合了多平台测序方法,重点关注长读长数据,以研究10个因高L1活性而被选中的人类肿瘤中的体细胞逆转录转座。我们开发了一种生物信息学算法,以核苷酸水平的分辨率揭示逆转录转座子介导的重排的模式和机制。此外,我们实施了时间估算方法来描述这些重排在肿瘤发生过程中的节奏。
在这10个肿瘤中,我们鉴定出6418个体细胞L1逆转录转座事件。其中,152个插入事件连接了基因组重排。尽管缺失仍然是主要类别,但易位成为第二常见的类型,约占这些结构变异形式的30%。值得注意的是,大约一半介导染色体间连接的逆转录转座子桥参与了相互易位——即非同源染色体之间的平衡交换,这在早期的短读长研究中基本无法检测到。长读长测序能够重建插入片段的内部结构,并揭示这些相互易位最常见的情况是涉及两个不同的L1分子,形成两条衍生染色体的连接点。值得注意的是,我们没有发现预先存在的体细胞L1插入之间发生整合后同源重组的证据。相反,我们的数据支持一个模型,即两个独立的L1插入被同时解析以产生重排。类似的原则也是相互倒位和更复杂的结构变异形成的基础,在这些变异中,独立的逆转录转座事件介导了串联重排中的多个连接点。
时间分析显示,约65%的L1插入事件发生在肿瘤演化的早期。在一个独立的、逆转录转座率较低的肿瘤队列中进行的比较分析证实了逆转录转座、TP53改变和全基因组DNA低甲基化之间的关联,同时进一步提示范可尼贫血通路的破坏与重排负荷升高有关。
总之,我们的观察结果证实体细胞L1活性是肿瘤基因组可塑性的一个主要贡献者,其影响远远超出了经典的插入突变。在这项工作中鉴定出的逆转录转座子介导的重排中,约有四分之三通过短读长测序无法检测到或被错误分类,这突显了这类结构变异在很大程度上被忽视了。总的来说,我们的发现将L1活性定位为人类癌症中大规模染色体不稳定性的早期驱动因素,大约每40到60个体细胞插入中就有1个会引起基因组重排。
LINE-1 (L1) 逆转录转座在人类癌症中产生体细胞基因组变异,但短读长测序限制了我们对其结构后果和动力学的理解。使用长读长测序,我们分析了10个具有异常高逆转录转座活性的肿瘤,包含超过6000个体细胞事件。我们发现L1介导的相互易位频繁发生,通常由两个同时发生在非同源染色体上的L1逆转录转座事件驱动。使用一个涵盖从低到高L1活性的独立肿瘤队列,我们估计逆转录转座子介导的重排的发生频率为每60个体细胞逆转录转座事件中出现1次。分子时间分析表明,这些事件发生在肿瘤发生的早期,确立了L1活性作为染色体不稳定性的早期驱动因素。我们的发现证明,在某些肿瘤中,L1对癌症基因组演化有实质性贡献。
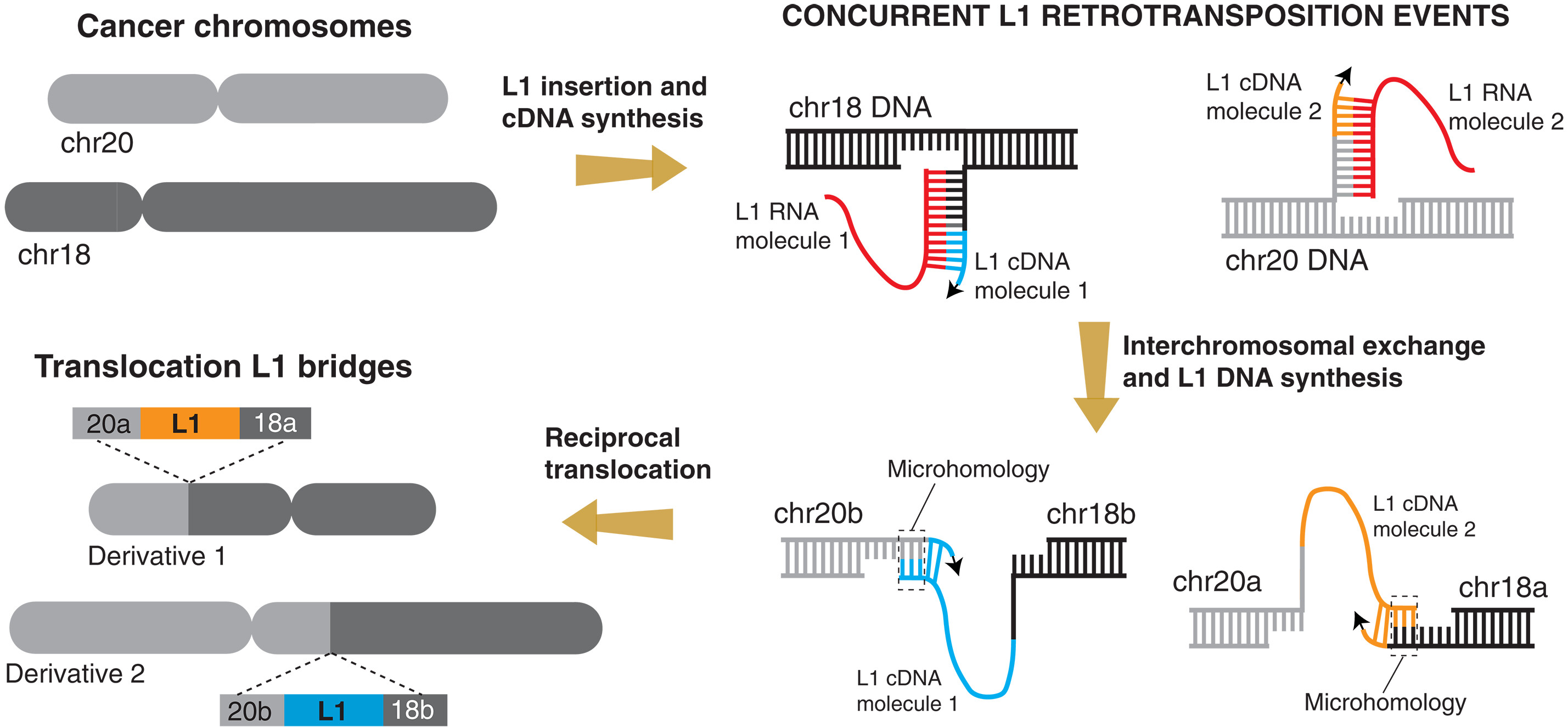
图:L1驱动染色体交换的模型示意图。在逆转录转座活性增强的肿瘤中,两个独立且同时发生的体细胞L1插入事件桥接了染色体间的断裂点,从而产生相互易位。每个插入事件起始于一条染色体,但其第二链的合成通过短序列微同源介导,在非同源染色体上完成。这一过程产生两个L1桥接结构,形成相互易位中两条衍生染色体的连接点。
Concurrent L1 retrotransposition events promote reciprocal translocations in human tumorigenesis