Cancer is a genetic disease, and much cancer research is focused on identifying carcinogenic mutations and determining how they relate to disease progression. Three papers demonstrate how mutations are processed through networks of protein interactions to promote cancer (see the Perspective by Cheng and Jackson). Swaney et al. focus on head and neck cancer and identify cancer-enriched interactions, demonstrating how point mutant–dependent interactions of PIK3CA, a kinase frequently mutated in human cancers, are predictive of drug response. Kim et al. focus on breast cancer and identify two proteins functionally connected to the tumor-suppressor gene BRCA1 and two proteins that regulate PIK3CA. Zheng et al. developed a statistical model that identifies protein networks that are under mutation pressure across different cancer types, including a complex bringing together PIK3CA with actomyosin proteins. These papers provide a resource that will be helpful in interpreting cancer genomic data. —VV
Genome-sequencing efforts over the past decade have profiled the genetic landscape of thousands of tumors and solidified the concept of cancer as a highly heterogeneous disease. Evidence from these efforts has revealed that thousands of genes are altered in cancer, presenting a high degree of complexity that can be challenging to translate into a molecular or clinical understanding. For example, head and neck squamous cell carcinoma (HNSCC) is the sixth most common malignancy worldwide and, despite a wealth of data detailing the genetic alterations in this tumor type, few targeted therapies are available. Therefore, HNSCC presents an opportunity to apply network biology approaches to identify new therapeutic targets and further our understanding of existing ones.
Network biology approaches have been successfully applied to bridge the gap between genetic alterations and clinical outcomes; however, such approaches rely heavily upon existing public databases of molecular interactions. With the growing recognition that molecular interactions can vary substantially across cellular contexts, the generation of networks in a cancer context represents a critical approach to interpreting and predicting cancer biology and its clinical outcomes.
To characterize the protein-protein interaction (PPI) landscape of HNSCC, we selected proteins based on altered molecular pathways identified from The Cancer Genome Atlas analysis of HNSCC tumors. Additional proteins were added based on genes with recurrent point mutations or a previously published association with HNSCC. PIK3CA (the gene encoding the alpha catalytic subunit of phosphoinositide 3-kinase) is the most commonly mutated oncogene in HNSCC, and although a few canonical mutations are well studied, there are many noncanonical mutations that are less well understood. We conducted affinity purification–mass spectrometry (AP-MS) analysis across three cell lines for 31 genes frequently altered in HNSCC, as well as 16 PIK3CA mutations. Two of the lines were HNSCC cell lines with RNA profiles representative of HNSCC patients, and one was an esophageal, non-tumerogenic cell line.
This network analysis uncovered 771 interactions from cancer and noncancerous cell states including wild-type and mutant protein isoforms. We found that 84% of these interactions had not been previously reported in public databases, providing a rich resource of new interactions with cancer relevance.
The data reveal a previously unidentified association of the fibroblast growth factor receptor (FGFR) tyrosine kinase 3 with Daple, a guanine-nucleotide exchange factor, resulting in activation of Gαi and PAK1/2 to promote cancer cell migration. This signaling pathway and cell migration can be effectively inhibited by FGFR inhibitors.
Furthermore, AP-MS analysis for 16 PIK3CA mutations revealed differences in PPIs. PIK3CA helical domain mutants, the most common in HNSCC, preferentially interact with the HER3 receptor tyrosine kinase. Analysis of isogenic xenografts in mice revealed that PIK3CA mutant interaction specificity can determine the in vivo response to HER3 inhibitors, with PIK3CA helical domain mutations conferring sensitivity to HER3 inhibitor treatment with CDX3379 and the H1047R kinase domain mutation conferring resistance.
We outline a framework for elucidating tumor genetic complexity through multidimensional PPI maps. This framework is applied to enhance our understanding of HNSCC and also breast cancer (see Kim et al., this issue). These works also suggest that a vast network of PPIs are left to be discovered. Such interactions, especially when combined with datatypes in a hierarchical model (see Zheng et al., this issue), can reveal new mechanisms of cancer pathogenesis, instruct the selection of therapeutic targets, and inform which point mutations in the tumor are most likely to respond to treatment. We anticipate that the application of this framework will be valuable to translating genetic alterations into a molecular and clinical understanding of the underlying biology of many diseases.
We outline a framework for elucidating tumor genetic complexity through multidimensional protein-protein interaction maps and apply it to enhancing our understanding of head and neck squamous cell carcinoma. This network uncovers 771 interactions from cancer and noncancerous cell states, including WT and mutant protein isoforms. Prioritization of cancer-enriched interactions reveals a previously unidentified association of the fibroblast growth factor receptor tyrosine kinase 3 with Daple, a guanine-nucleotide exchange factor, resulting in activation of Gαi- and p21-activated protein kinase 1/2 to promote cancer cell migration. Additionally, we observe mutation-enriched interactions between the human epidermal growth factor receptor 3 (HER3) receptor tyrosine kinase and PIK3CA (the alpha catalytic subunit of phosphatidylinositol 3-kinase) that can inform the response to HER3 inhibition in vivo. We anticipate that the application of this framework will be valuable for translating genetic alterations into a molecular and clinical understanding of the underlying biology of many disease areas.
癌症是一种基因疾病,大量癌症研究聚焦于识别致癌突变并确定它们与疾病进展的关系。三篇论文展示了突变如何通过蛋白质相互作用网络被处理从而促进癌症(参见Cheng与Jackson的观点文章)。Swaney等人专注于头颈癌,识别出癌症富集的相互作用,证明了PIK3CA(一种在人类癌症中频繁突变的激酶)的点突变依赖性相互作用如何预测药物反应。Kim等人专注于乳腺癌,识别出两个与肿瘤抑制基因BRCA1功能相连的蛋白质,以及两个调控PIK3CA的蛋白质。Zheng等人开发了一个统计模型,用于识别在不同癌症类型中承受突变压力的蛋白质网络,包括一个将PIK3CA与肌动球蛋白复合物聚集在一起的复合体。这些论文提供了有助于解读癌症基因组数据的资源。—VV
过去十年的基因组测序工作描绘了数千种肿瘤的基因图谱,并巩固了癌症作为一种高度异质性疾病的概念。这些工作获得的证据表明,数千个基因在癌症中发生改变,呈现出高度的复杂性,这给转化为分子或临床理解带来了挑战。例如,头颈部鳞状细胞癌是全球第六大常见恶性肿瘤,尽管有大量数据详细描述了此类肿瘤的基因改变,但可用的靶向疗法却很少。因此,头颈鳞癌为应用网络生物学方法识别新治疗靶点并深化对现有靶点的理解提供了机会。
网络生物学方法已成功应用于弥合基因改变与临床结果之间的鸿沟;然而,这类方法在很大程度上依赖于现有的分子相互作用公共数据库。随着人们越来越认识到分子相互作用在不同细胞背景下可能存在显著差异,在癌症背景下构建网络成为解读和预测癌症生物学及其临床结果的关键方法。
为了描绘头颈鳞癌的蛋白质-蛋白质相互作用景观,我们根据头颈鳞癌肿瘤的癌症基因组图谱分析所识别的改变分子通路来筛选蛋白质。另外,基于具有复发性点突变的基因或先前发表的与头颈鳞癌相关的基因,我们添加了其他蛋白质。PIK3CA(编码磷脂酰肌醇3-激酶的α催化亚基的基因)是头颈鳞癌中最常发生突变的癌基因,尽管少数经典突变已得到充分研究,但许多非经典突变尚不明确。我们对头颈鳞癌中频繁改变的31个基因以及16个PIK3CA突变,在三种细胞系中进行了亲和纯化-质谱分析。其中两个是头颈鳞癌细胞系,其RNA谱能代表头颈鳞癌患者,另一个是食管非致瘤性细胞系。
该网络分析揭示了来自癌症和非癌细胞状态(包括野生型和突变型蛋白质亚型)的771种相互作用。我们发现其中84%的相互作用在公共数据库中未曾报道过,这为癌症相关的新相互作用提供了丰富的资源。
数据揭示了一个先前未识别的成纤维细胞生长因子受体3酪氨酸激酶与Daple(一种鸟嘌呤核苷酸交换因子)的关联,导致Gαi和PAK1/2的激活,从而促进癌细胞迁移。该信号通路和细胞迁移可被FGFR抑制剂有效抑制。
此外,对16个PIK3CA突变的AP-MS分析揭示了蛋白质-蛋白质相互作用的差异。PIK3CA螺旋结构域突变(头颈鳞癌中最常见的类型)优先与HER3受体酪氨酸激酶相互作用。对小鼠等基因异种移植瘤的分析表明,PIK3CA突变体的相互作用特异性可以决定体内对HER3抑制剂的反应,PIK3CA螺旋结构域突变对CDX3379这种HER3抑制剂治疗敏感,而H1047R激酶结构域突变则产生耐药性。
我们概述了一个通过多维蛋白质-蛋白质相互作用图谱来阐明肿瘤基因复杂性的框架。该框架用于深化我们对头颈鳞癌以及乳腺癌的理解(参见本期Kim等人的文章)。这些工作还表明,仍有大量的蛋白质-蛋白质相互作用网络有待发现。这些相互作用,尤其是在分层模型中与其他数据类型结合时(参见本期Zheng等人的文章),可以揭示癌症发病机制的新机制,指导治疗靶点的选择,并提示肿瘤中的哪些点突变最有可能对治疗产生反应。我们预计,应用此框架将有助于将基因改变转化为对多种疾病潜在生物学的分子和临床理解。
我们概述了一个通过多维蛋白质-蛋白质相互作用图谱来阐明肿瘤基因复杂性的框架,并将其应用于深化对头颈部鳞状细胞癌的理解。该网络揭示了来自癌症和非癌细胞状态(包括野生型和突变型蛋白质亚型)的771种相互作用。对癌症富集相互作用的优先排序揭示了一个先前未识别的成纤维细胞生长因子受体3酪氨酸激酶与Daple(一种鸟嘌呤核苷酸交换因子)的关联,导致Gαi和p21活化激酶1/2的激活,从而促进癌细胞迁移。此外,我们观察到人表皮生长因子受体3酪氨酸激酶与PIK3CA(磷脂酰肌醇3-激酶的α催化亚基)之间存在突变富集的相互作用,这可以提示体内对HER3抑制的反应。我们预计,应用此框架将有助于将基因改变转化为对许多疾病领域潜在生物学的分子和临床理解。
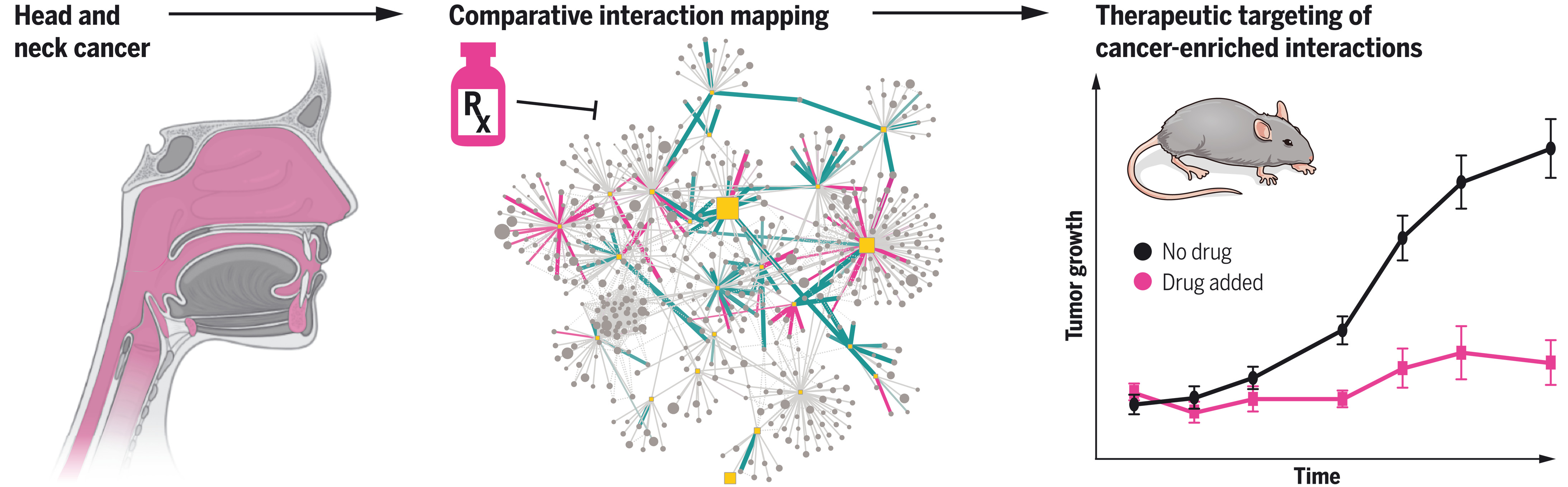
图:头颈鳞状细胞癌相互作用组分析。对头颈鳞状细胞癌中常见变异基因的蛋白质-蛋白质相互作用分析揭示了可作为治疗性靶点以抑制肿瘤生长的相互作用。
A protein network map of head and neck cancer reveals PIK3CA mutant drug sensitivity