The heterogeneity of mammalian tumors has been well documented, but it remains unknown how differences between individual cells lead to metastasis and spread throughout the body. Quinn et al. created a Cas9-based lineage tracer and used single-cell sequencing to generate phylogenies and follow the movement of metastatic human cancer cells implanted in the lung of a mouse xenograph model. Using this model, they found that within the same cell line, cancer cells exhibited diverse metastatic phenotypes. These subclones exhibited differential gene expression profiles, some of which were previously associated with metastasis.
Cancer progression is fundamentally an evolutionary process, involving multiple distinct steps from oncogenic transformation to metastatic dissemination to development of therapeutic resistance. Defining the timing and molecular nature of each step in this process is critical to understanding cancer biology and to devising effective therapeutic strategies. Yet, it is challenging to directly observe these events because they unfold stochastically, in rare subpopulations of cells, and over long time scales.
By reconstructing the phylogenetic relationships of tumor cells, it is possible to infer important features of these abstruse, past events, such as the timing or directionality of metastatic dissemination. Advances in Cas9-based molecular recording technologies have now enabled the high-resolution mapping of subclonal cellular lineages that, when paired with single-cell RNA sequencing, can be used to infer distant events in a cell’s past history and connect them to its present state.
We refined and applied our Cas9-based lineage recorder to study metastatic dissemination in implanted tumor cells in a mouse. These advances allowed us to resolve deep and accurate single-cell lineages for tens of thousands of metastatically disseminated cancer cells in a mouse over 2 months of growth.
We leveraged the reconstructed phylogenetic trees to measure the frequency of metastatic dissemination at single-cell resolution. Unexpectedly, we found that tumors arising from individual implanted cancer cells showed dramatic differences in their capacity to spread to distant tissues, ranging from completely nonmetastatic (i.e., never leaving the primary site) to aggressively metastatic (i.e., frequently transiting between tissues). We paired this high-resolution lineage information with single-cell RNA sequencing to reveal characteristic sets of genes that underlie these divergent metastatic phenotypes. This analysis identified some genes that are known to modulate metastatic phenotype in lung cancers (such as ID3 and REG4) and others that were unexpected (such as KRT17). Using CRISPR-based perturbations in two distinct lung cancer cell lines, we validated that over- or underexpression of genes identified by this analysis was sufficient to modulate cancer invasion phenotypes.
We then showed that the diversity of metastatic phenotypes observed in vivo stemmed from preexisting, heritable cellular states. Specifically, the heterogeneity in the expression of metastasis-associated genes existed before implantation into the mice, and cells with higher metastatic transcriptional signatures preimplantation went on to be more metastatic in vivo. Moreover, when cells from the same clone were implanted into different mice, they exhibited nearly identical metastatic behavior.
Finally, we traced the complex and multidirectional routes through which the metastases disseminated from tissue to tissue and identified the mediastinal lymphatic tissue as a transit hub for metastasis. We also found numerous examples of different seeding topologies, including reseeding (i.e., cancer cells returning to the primary tumor site from a metastasis) and seeding cascades (multistep metastatic seeding).
High-resolution lineage recording of tumor cells revealed a rich diversity of metastatic phenotypes and behaviors that would have otherwise been unobservable. This illuminated aspects of cancer biology that are essential to understanding disease progression, such as the rates, tissue routes, and transcriptional drivers of metastasis. We anticipate that our approach will be broadly applicable to future studies of other aspects of cancer biology, such as the evolution of drug resistance, and, more generally, of biological processes that unfold over many cell generations.
Detailed phylogenies of tumor populations can recount the history and chronology of critical events during cancer progression, such as metastatic dissemination. We applied a Cas9-based, single-cell lineage tracer to study the rates, routes, and drivers of metastasis in a lung cancer xenograft mouse model. We report deeply resolved phylogenies for tens of thousands of cancer cells traced over months of growth and dissemination. This revealed stark heterogeneity in metastatic capacity, arising from preexisting and heritable differences in gene expression. We demonstrate that these identified genes can drive invasiveness and uncovered an unanticipated suppressive role for KRT17. We also show that metastases disseminated via multidirectional tissue routes and complex seeding topologies. Overall, we demonstrate the power of tracing cancer progression at subclonal resolution and vast scale.
哺乳动物肿瘤的异质性已有充分记录,但个体细胞间的差异如何导致转移并在全身扩散仍属未知。Quinn等人创建了一种基于Cas9的谱系追踪技术,并利用单细胞测序来生成系统发育树,追踪植入小鼠异种移植模型肺部中的转移性人类癌细胞的移动。利用该模型,他们发现同一细胞系内的癌细胞表现出多样化的转移表型。这些亚克隆展现出差异化的基因表达谱,其中一些此前已被认为与转移相关。
癌症进展从根本上说是一个进化过程,涉及从致癌转化到转移扩散再到产生治疗耐药性的多个不同步骤。界定这一过程中每个步骤的时间点和分子本质,对于理解癌症生物学和制定有效的治疗策略至关重要。然而,直接观察这些事件具有挑战性,因为它们随机发生、出现在稀少的细胞亚群中,并且跨越很长时间尺度。
通过重建肿瘤细胞的系统发育关系,可以推断这些深奥过去事件的重要特征,例如转移扩散的时间点或方向性。基于Cas9的分子记录技术的进步,现已能够对亚克隆细胞谱系进行高分辨率图谱绘制。当与单细胞RNA测序结合时,可用于推断细胞过去历史中的遥远事件,并将其与细胞当前状态联系起来。
我们改进并应用了基于Cas9的谱系记录器来研究植入小鼠体内的肿瘤细胞的转移扩散。这些进步使我们能够在2个多月的生长过程中,解析出数万个已发生转移扩散的癌细胞深度且准确的单细胞谱系。
我们利用重建的系统发育树,以单细胞分辨率测量了转移扩散的频率。出乎意料的是,我们发现由单个植入癌细胞产生的肿瘤在向远处组织扩散的能力上存在巨大差异,范围从完全非转移性(即从未离开原发部位)到高度侵袭性转移性(即频繁在组织间迁移)。我们将这种高分辨率谱系信息与单细胞RNA测序相结合,揭示了构成这些不同转移表型的特征性基因集合。该分析识别出一些已知能调节肺癌转移表型的基因(如ID3和REG4)以及其他意外发现的基因(如KRT17)。通过在两种不同的肺癌细胞系中进行基于CRISPR的干扰实验,我们验证了通过此分析识别出的基因的过表达或表达不足足以调节癌症侵袭表型。
随后,我们表明在体内观察到的转移表型多样性源于预先存在的、可遗传的细胞状态。具体来说,转移相关基因表达的异质性在植入小鼠之前就已存在,且植入前具有较高转移转录特征的细胞在体内会表现出更强的转移性。此外,当同一克隆的细胞被植入不同小鼠体内时,它们表现出几乎完全相同的转移行为。
最后,我们追踪了转移灶在组织间扩散的复杂且多方向的路径,并确定纵隔淋巴组织是转移的中转枢纽。我们还发现了许多不同的播种拓扑结构实例,包括再播种(即癌细胞从转移灶返回原发肿瘤部位)和级联播种(多步骤的转移性播种)。
肿瘤细胞的高分辨率谱系记录揭示了过去无法观察到的丰富多样的转移表型和行为。这阐明了对于理解疾病进展至关重要的癌症生物学方面,例如转移的速率、组织路径和转录驱动因素。我们预计我们的方法将广泛适用于未来对癌症生物学其他方面的研究,例如耐药性的演变,以及更广泛地,适用于跨越多个细胞世代发生的生物过程研究。
肿瘤群体的详细系统发育可以重述癌症进展过程中关键事件的历史和时间顺序,例如转移扩散。我们应用了一种基于Cas9的单细胞谱系追踪技术来研究肺癌异种移植小鼠模型中转移的速率、路径和驱动因素。我们报告了在数月生长和扩散过程中追踪到的数万个癌细胞的深度解析系统发育树。这揭示了转移能力的显著异质性,其源于基因表达上预先存在且可遗传的差异。我们证明了这些被识别出的基因能够驱动侵袭性,并揭示了KRT17一个未曾预料到的抑制性作用。我们还表明转移是通过多方向的组织路径和复杂的播种拓扑结构进行扩散的。总体而言,我们展示了在亚克隆分辨率和巨大规模上追踪癌症进展的强大能力。
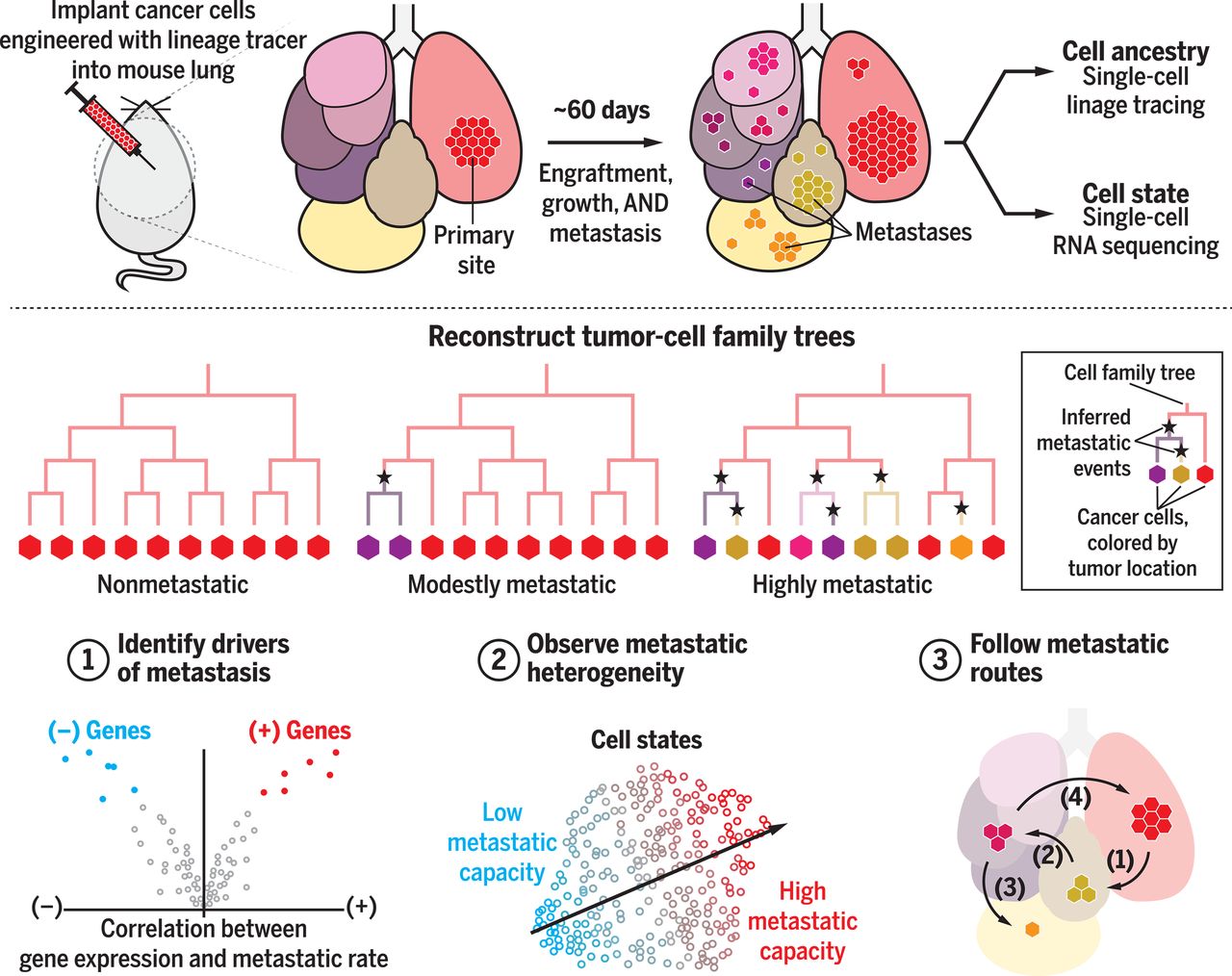
图:胰腺癌中的隐秘抗原。非编码基因的异常翻译会产生非常规(隐秘)肽段,这些肽段能够在胰腺癌中被 HLA-I 处理并呈递。隐秘肽段具有很强的免疫原性,相应的 TCR 能够在体外和体内识别并杀死患者来源的胰腺癌细胞团。nuORF,新未注释的开放阅读框;dORF,下游开放阅读框;Ribo-seq,核糖体测序;CTLs,细胞毒性 T 淋巴细胞;IFN-γ,干扰素-γ;TNFα,肿瘤坏死因子-α。
Single-cell lineages reveal the rates, routes, and drivers of metastasis in cancer xenografts